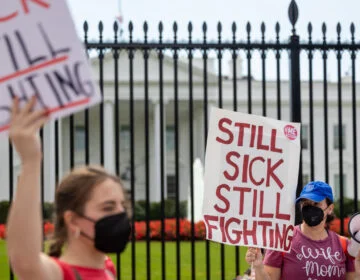Want to know which COVID variant you have? Rutgers scientists have a test for that
A new PCR test could more quickly and easily identify COVID variants.

(Courtesy of the Tyagi lab at Rutgers University)
The COVID-19 pandemic has changed everything. What should we know about how you approach the world now? How has the pandemic changed your social life, your work life, your interactions with your neighbors? Get in touch here.
Tests that can detect whether someone is infected with the virus that causes COVID-19 are now readily available, but those tests can’t determine which variant of the coronavirus a person has.
Rutgers University scientists say it could be a possibility in the future, however. They’ve developed a PCR test that identifies COVID-19 variants — which, they say, would help epidemiologists track the disease, as well as physicians better treat patients.
Scientists worldwide already track variants to identify which ones are predominant in a particular region, and to keep an eye on new mutations. But the process involves complicated genome sequencing led by microbiologists who evaluate select patient samples.
The Rutgers test, on the other hand, could be used in concert with any COVID-19 PCR (polymerase chain reaction) test, yielding information at a faster rate and at a larger scale, the university’s scientists say.
The test works by using molecular beacons, which create a signal when a specific variant of the virus is present. Secondary to an initial COVID-19 test, it would be used on COVID-positive samples, in either a lab or hospital setting, to determine which variant a person is infected with. In the hospital setting, it could render results in a few hours, just like any other PCR test on the market, the Rutgers scientists say, compared to sequencing, which has a lag of a couple of weeks.
Details about the test were recently published in the Journal of Molecular Diagnostics. The
Rutgers scientists are currently working to make use of the test more widely, and, eventually, to apply for Emergency Use Authorization from the Food and Drug Administration in order to commercialize it.
Rutgers scientists currently are working to get FDA-approval for and commercialize the test, but they don’t know how long it could take before it goes to market.
The study’s first author Ryan Dikdan, a doctoral student at the university, said he and his colleagues want to track variants as they emerge in a cost-effective way. They say the test would cost the same as currently available PCR tests, and wouldn’t require any additional equipment, time, or expertise to run.
“Since this is just a normal PCR test, it’s scalable, and you can run it on every single positive COVID sample in order to track emerging variants, in order to track whether a new variant such as Omicron has made its way to America or not. And that’s what we showed in the paper, which was that we picked up when Omicron was coming over,” Dikdan said.
Some COVID treatments are not ‘one size fits all’
The test could also help physicians when they prescribe monoclonal antibody treatments, he said. Monoclonal antibodies are given to a small percentage of COVID-19 patients, of whom are at high-risk of serious illness, such as transplant patients or people with immunosuppressive conditions.
But there are very few monoclonal antibody treatments available, and each treatment only targets specific variants of the coronavirus. Physicians currently try to prescribe those that are most likely to be effective, said Dr. Thomas Fekete, a professor of medicine, microbiology and immunology at the Lewis Katz School of Medicine at Temple University.
“The goal is to match up the very specific configuration of the antibody with that of the virus … And if you have an outdated antibody that doesn’t recognize some new mutations, it’s like giving nothing,” he said.
Fekete likened how monoclonal antibody treatments aren’t one size fits all to using someone else’s car key on your vehicle.
“In a sense, this is a little bit of what we’re trying to figure out,” he said. “Even small differences in the keys may mean that your car will not unlock. So, we’re trying to get it to the point where we can do it effectively and quickly.”
Knowing which variant of COVID-19 a patient is infected with would help target monoclonal antibody treatment faster and more effectively, Dikdan said of the Rutgers test.
“The idea of being able to check which subvariant they have by using a PCR test, which is very quick compared to sequencing or anything else, could improve patient outcomes. It would not be giving a drug that won’t do anything,” he said.
Fekete pointed out that the number of patients who are prescribed monoclonal antibody treatment is small, and that the majority of healthy people don’t need to know what variant of COVID-19 they have.
“It’s a little bit of a boutique item what they’re describing, and it could be very helpful in tracking the pandemic and giving us a better sense of what’s happening. But it’s not going to replace the things that you and I are going to be getting tested for tomorrow or the next day,” he said.
But from a population perspective; “If I’m a public health expert, or the CDC, or some other agency, I would say I really want to know everything about this virus,” Fekete said.
Dikdan also said that if resistance to newer antiviral treatments ever were to arise, the test could inform decision-making around treatment.
Improved variant tracking could help curb the pandemic
In addition to understanding how effective monoclonal antibodies will be, there are a variety of reasons to track variants, said Dr. Frederick Bushman, a professor and chair of microbiology at the University of Pennsylvania’s Perelman School of Medicine.
Variants have different degrees of severity and transmissibility, and vaccine effectiveness, too, can vary depending on the variant. Scientists can also use information about variants to understand the trajectory of viral evolution, Bushman said. The Bushman lab has sequenced about 6,000 genomes of the virus in the Delaware Valley, which has helped scientists and health officials understand more about the trajectory of the pandemic.
“If we know what variants are involved, what their properties are, we can tell what to expect by sort of forecasting by the experience of other places that are ahead of us, based on our sequence and knowledge of the different variants,” Bushman said.
Scientists also are tracking variants among animals. For example, researchers in Canada found a population of deer infected with an Omicron-like variant of coronavirus, and one human who had close contact with deer also was infected with the variant.
“What if something changes in SARS-CoV-2 so they can jump into humans? All of this is tracked by the deep sequencing methods,” Bushman explained.
He said it’s difficult to predict what the future holds for the COVID-19 pandemic. However, he surmised that the virus will continue to evolve, and will become less pathogenic in order to spread more efficiently. Cases of COVID-19 are rising again, and more infectious variants of Omicron have been detected.
“The good news is that the virus seems to be less pathogenic, but with more and more people getting infected, there’s sort of a denominator effect. There’ll be some people who are immunocompromised, very vulnerable, and get severely ill,” Bushman said.
“So, there’s not much downside to wearing masks. If you’re not vaccinated, you should definitely get vaccinated. We’re just going to have to come to grips with the fact that the numbers of infections do seem to be starting up. We need to be thinking, ‘How are we going to react to aggressive infection by a less pathogenic virus?’”
WHYY is your source for fact-based, in-depth journalism and information. As a nonprofit organization, we rely on financial support from readers like you. Please give today.




![CoronavirusPandemic_1024x512[1]](https://whyy.org/wp-content/uploads/2020/03/CoronavirusPandemic_1024x5121-300x150.jpg)